This is My Jam has a simple dataset for our purposes. The very first Hadoop code of mine will try to extra the number of likes a jam gets. We will not order them for now.
Every Hadoop code I develop has a similar pattern. First, I write the code and its unit test simultaneously. I compile them and run the unit test immediately. In the second step, I code a driver class. The aim of this is to run the application locally, without the worry to scale. This will help me to simulate my Hadoop application as if it runs on a cluster consisting of one node running both the mapper and the reducer. As the last step, I deploy the application to the RPi Hadoop cluster and analyse its performance characteristics. I will try to stick this flow as much as possible.
Each record of the likes.tsv file consists of userID and the jamID she liked, separated by a tab character. The record itself is unique. Our output will be a list of the jamIDs and how many different users liked that. We won’t introduce an explicit ordering to the output. Therefore it will be ordered by jamIDs in a descending fashion.
Implementing mapreduce applications is divided into two related and consecutive steps. The first one is mapping the input to an intermediate state. This step is not an arbitrary form, rather it should comply with the input structure of the reducer, which is the second step. So we will be talking about a mapper class and a reducer class. Here is the mapper:
public static class TokenizerMapper
extends Mapper<LongWritable, Text, Text, IntWritable>{
private final static IntWritable one = new IntWritable(1);
public void map(LongWritable key, Text value, Context context
) throws IOException, InterruptedException {
String[] userAndJam = value.toString().split("\t");
context.write(new Text(userAndJam[1]), one);
}
}
We split a record into two parts and use only the jamID. Actually, the userID is also used, not directly but to increase the counter for the jamID. It adds up to that counter by one.
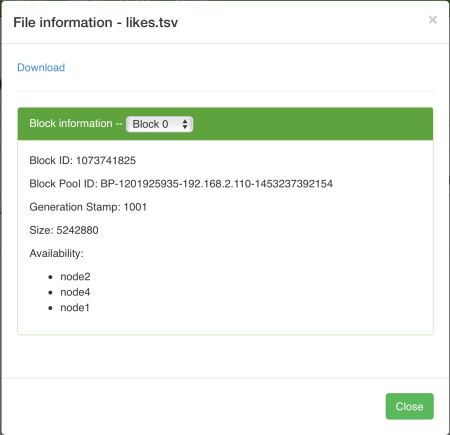
So how is this code used? Actually, the whole input file is divided into mostly equal sized parts. Each part is fed into a datanode. The datanode runs this mapper application and produces an output. In our case, the output is a key value pair, where key is the jamID and value is an integer, in fact simply a 1.
As I said before, the mapreduce framework assigns datanodes a part of the input data. When a datanode finishes its map task, the mapreduce framework gets its output and merges it with the other outputs. Since they all have a key, the framework simply appends the values for the same keys. In our case, node1 may generate the following map output:
jam1 1 jam2 1 jam1 1 jam3 1 jam1 1
The node2 may generate an entirely different map output as in:
jam2 1 jam2 1 jam1 1 jam4 1
The intermediate output of all the map tasks, which will be the input for the reducer, will be:
jam1 [1, 1, 1, 1] jam2 [1, 1, 1] jam3 [1] jam4 [1]
Note that, the intermediate output is sorted by keys. The same ordering will be preserved in the output of the reducer, unless an explicit sorting mechanism is implemented in the reducer.
However, the order of the values is NOT preserved or even deterministic. I emphasised it by mixing the order from different nodes. node1 is represented by blue and node2 is represented by red.
With that input, what the reducer should do becomes very trivial. It simply adds 1s in the list for each jamID. The result will be:
jam1 4 jam2 3 jam3 1 jam4 1
Note that the order of the output is the same as the order in the intermediate step. Here is the code for the reducer:
public static class IntSumReducer
extends Reducer<Text,IntWritable,Text,IntWritable> {
private IntWritable result = new IntWritable();
public void reduce(Text key, Iterable<IntWritable> values,
Context context
) throws IOException, InterruptedException {
int sum = 0;
for (IntWritable val : values) {
sum += val.get();
}
result.set(sum);
context.write(key, result);
}
}
To bring all those together and encapsulate as a class, I modified the tutorial code and add it to my github repository.
I preferred to use MRUnit for unit tests. For any mapreduce class, we can generate three units tests:
- Unit test for mapper: This is for testing the mapper only
- Unit test for reducer: This is for testing the reducer only
- Unit test for mapper-reducer: This is the complete test from start to end
@Test
public void testMapper() {
mapDriver.withInput(new LongWritable(), new Text(
"c1066039fa61eede113878259c1222d1\t5d2bc46196d7903a5580f0dbedc09610"));
mapDriver.withOutput(new Text("5d2bc46196d7903a5580f0dbedc09610"), new IntWritable(1));
try {
mapDriver.runTest();
} catch (Exception e) {
e.printStackTrace();
}
}
@Test
public void testReducer() {
List<IntWritable> values = new ArrayList<IntWritable>();
values.add(new IntWritable(1));
values.add(new IntWritable(1));
reduceDriver.withInput(new Text("5d2bc46196d7903a5580f0dbedc09610"), values);
reduceDriver.withOutput(new Text("5d2bc46196d7903a5580f0dbedc09610"), new IntWritable(2));
try {
reduceDriver.runTest();
} catch (Exception e) {
e.printStackTrace();
}
}
I only tested one record for mapper and reducer. For the full test, I decided to work on a bigger (3!) data set:
@Test
public void testMapReduce() {
List<Pair<LongWritable, Text>> inputLines = new ArrayList<Pair<LongWritable, Text>>(3);
Pair<LongWritable, Text> i1 = new Pair<LongWritable, Text>(new LongWritable(1), new Text(
"c1066039fa61eede113878259c1222d1\t5d2bc46196d7903a5580f0dbedc09610"));
Pair<LongWritable, Text> i2 = new Pair<LongWritable, Text>(new LongWritable(2), new Text(
"c1066039fa61eede113878259c1222dl\t5d2bc46196d7903a5580f0dbedc09610"));
Pair<LongWritable, Text> i3 = new Pair<LongWritable, Text>(new LongWritable(3), new Text(
"c1066039fa61eede113878259c1222d1\t5d2bc46l96d7903a5580f0dbedc09610"));
inputLines.add(i1);
inputLines.add(i2);
inputLines.add(i3);
mapReduceDriver.withAll(inputLines);
List<Pair<Text, IntWritable>> outputLines = new ArrayList<Pair<Text, IntWritable>>(2);
Pair<Text, IntWritable> o1 = new Pair<Text, IntWritable>(new Text("5d2bc46196d7903a5580f0dbedc09610"), new IntWritable(2));
Pair<Text, IntWritable> o2 = new Pair<Text, IntWritable>(new Text("5d2bc46l96d7903a5580f0dbedc09610"), new IntWritable(1));
outputLines.add(o1);
outputLines.add(o2);
mapReduceDriver.withAllOutput(outputLines);
try {
mapReduceDriver.runTest();
} catch (Exception e) {
e.printStackTrace();
}
}
Although the tests differ from each other and concentrate on different aspects, the idea is the same. Add the input data, which represent the records. Add the expected output and runTest().
For the sake of completeness, I also uploaded the unit tests to my github repository.
After running unit tests, we can either run the application on the HDFS cluster (which I do NOT recommend) or run the application with a driver class. The importance of the driver class is that, it enables us to see its performance on the local file system. Furthermore, it gives us the opportunity to remove last splinters before the scalability comes in to the scene. The driver class I have written can again be found in the hub.
Before running the driver, however, we need to create a new Hadoop configuration. This is essential since the driver class will not have any access to the HDFS and the trackers. Instead, it will use the local mapred job tracker. We create the following configuration file, mapred-local.xml, under /usr/local/hadoop/etc/hadoop/ by copying from the mapred-site.xml file:
cp /usr/local/hadoop/etc/hadoop/mapred-site.xml /usr/local/hadoop/etc/hadoop/mapred-local.xml
<configuration>
<property>
<name>fs.default.name</name>
<value>file:///</value>
</property>
<property>
<name>mapred.job.tracker</name>
<value>local</value>
</property>
</configuration>
javac -cp ${HADOOP_JAVA_PATH} -d /usr/local/hadoop/my_jam_analysis/ /usr/local/hadoop/*.java
java -cp ${HADOOP_JAVA_PATH} org.junit.runner.JUnitCore tr.name.sualp.merter.hadoop.myjam.LikedJamCountMapperReducerTest
java -cp ${HADOOP_JAVA_PATH} tr.name.sualp.merter.hadoop.myjam.LikedJamCountDriver -conf /usr/local/hadoop/etc/hadoop/mapred-local.xml /media/sf_Downloads/This\ is\ My\ Jam\ Dataset/likes.tsv output
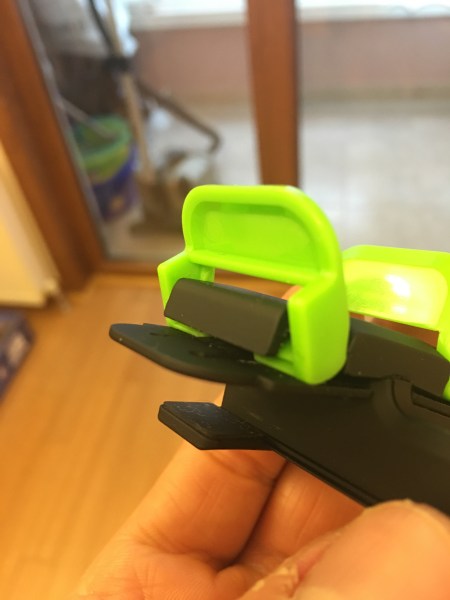
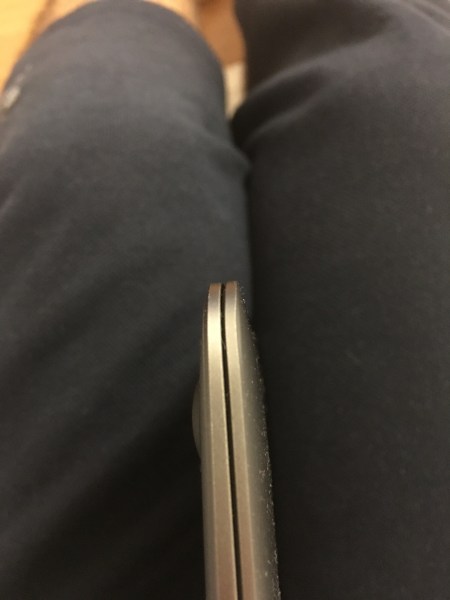





 That location may be the weakest construction point on the lid of Macbook, I do not know, but keep an eye for those kind of deficiencies. I lowered the height by using D in place of F but it does not hold as powerful as F. There is no such problem on the other side of the clip. My iPad and iPhone are ok.
That location may be the weakest construction point on the lid of Macbook, I do not know, but keep an eye for those kind of deficiencies. I lowered the height by using D in place of F but it does not hold as powerful as F. There is no such problem on the other side of the clip. My iPad and iPhone are ok.